Large Language Models as Information Extractors in Healthcare
Introduction
Doctors and hospitals rely on medical records to make life-or-death decisions. These records contain vital information such as diagnoses, treatments, medications, lab results, and clinical observations that guide patient care.
But medical records are long, complex, and often messy. Finding the important details can take hours of careful review, and mistakes or delays can affect patient outcomes. Accurate information extraction is not only critical for doctors and patients, but it also supports hospital operations, research, and reporting to agencies like the Centers for Medicare and Medicaid Services (CMS).
AI tools, like large language models (LLMs), can help speed up information extraction from medical records. But they have a problem: when given long and messy notes, they often get confused and miss the most important details. This limits how much hospitals can rely on AI for critical tasks. Retrieval-augmented generation (RAG) can help mitigate this challenge by reducing the amount of information providing to LLMs, however, this field is underexplored. Thus, we introduce a benchmark for evaluating and comparing retrieval methods for EHR information extraction. To help LLMs focus on the right information, we tested different retrieval strategies that guide the model to the most relevant parts of the records before it answers questions. Through this framework, researchers are able to compare different strategies and see which ones work best in medical settings.
We tested our methods using real patient records from the MIMIC-III database, which contains thousands of de-identified unstructured clinical notes from a hospital’s intensive care units. These notes come in many forms such as nursing notes, doctor’s observations, lab reports, and discharge summaries, and they vary in length and style.
This work provides
- A benchmark for evaluating retrieval approaches on medical information.
- A comparison of different retrieval strategies to see which methods work best.
- Insights for researchers on how to make LLMs more accurate and reliable in healthcare settings.
Methodology
To test how well LLMs can locate important clinical details, we created a “needle in a haystack” approach using real patient records. Our process is visually shown in Figure 1.
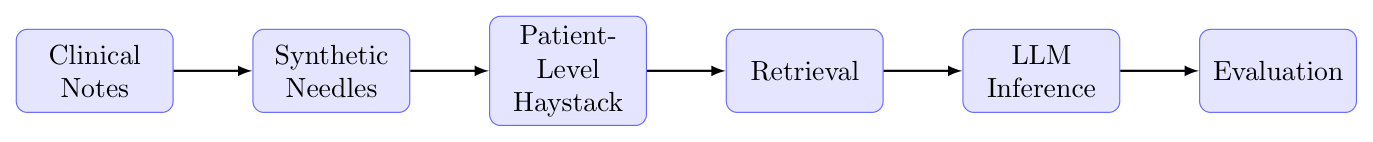
Data Preparation
We defined meaningful quries by using guidelines of the Severe Sepsis and Septic Shock Early Management Bundle (SEP-1) from the CMS specifications manual. We constructed these queries based on five elements described in the manual:
- Administrative Contraindication to Care–Severe Sepsis
- Directive for Comfort Care or Palliative Care–Severe Sepsis
- Clinical Trial–Severe Sepsis
- Severe Sepsis Present
- Vasopressor Administration–Severe Sepsis.
We then randomly genertated 100 example “needle” statements that provide positive responses to the queries and serve as ground truth evidence for evaluation. These needles were inserted randomly into each patient’s record, creating our “haystack” or a long document full of extra information with one important fact hidden inside. Our benchmark consists of 10,000 patients (461,290 total clinical notes) resulting in 10,000 haystacks used for evaluation.
Retrieval approaches
We wanted to determine which retrieval methods could help LLMs find the needles most effectively so we testd six approaches:
- BM25 – finds exact keyword matches. Works well if the question and text use the same words.
- FAISS – converts text into vectors and finds passages that are “similar in meaning.”
- FAISS + MMR – improves FAISS by choosing results that are diverse and non-redundant.
- Hybrid approach – combines BM25 and FAISS to incorporate meaning and exact matching.
- Semantic chunking – splits long notes into meaningful segments so the focus is on smaller, coherent chunks.
- SPLADE – an advanced model that matches semantically similar words even if the wording is different.
We also tested a baseline approach where the LLM received the entire patient record without retrieval. This allowed us to evaluate whether focusing on relevant passages improves performance.
LLM Prompting and Answer Generation
ALL LLM tests were run using AWS Bedrock, a secure platform to protect patient privacy. We used the DeepSeek-V3 model because of its favorable balance between cost and performance.
For each retrieval method, the most relevant passages were selected and provided to the LLM along with the query. The LLM then performed two tasks:
- Classification: Determine whether the retrieved passages contained enough information to answer the query. A positive “Yes” response indicated that the model identified relevant evidence.
- Extraction: Identify and extract the specific clinical evidence related to the query. A match between the extracted evidence and the synthetic needles indicated that the correct information had been found.
This way, we could measure both if the LLM could answer the query when given limited context and if the limited context even contained the corrected the correct information.
Evaluation
We evaluated model performance by comparing the LLM’s responses to the known synthetic needles inserted into each document. This allowed us to measure which retrieval strategies helped the LLM locate the hidden information most often and how accurately the model could identify and extract key clinical evidence
Results
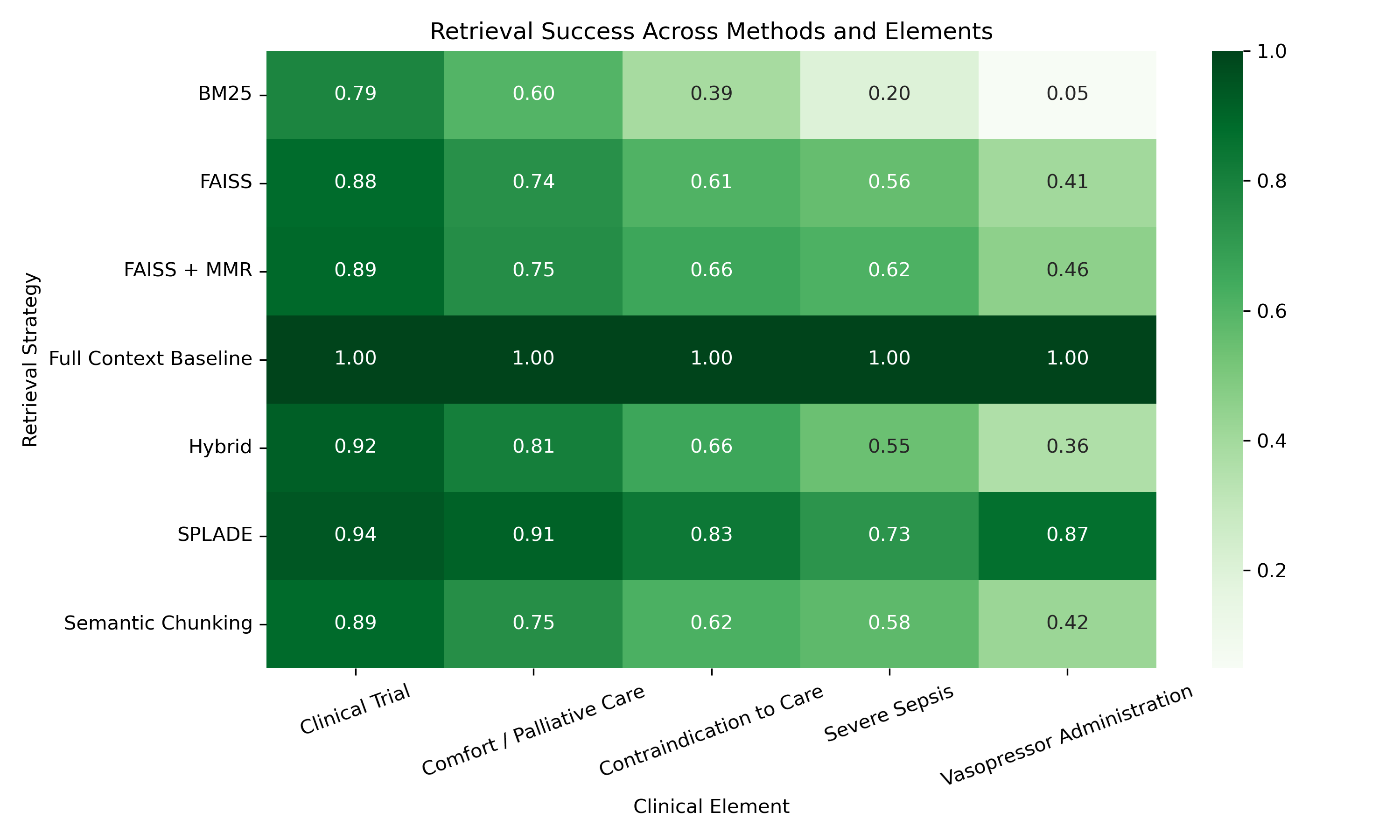
Our experiments revealed clear differences in how well each retrieval method helped the LLM find the hidden “needle” information within patient records. SPLADE consistently achieved the highest retrieval success, meaning it was the most effective at finding the relevant clinical evidence across all five clinical elements. In contrast, BM25 showed the lowest performance, struggling to retrieve relevant passages when the wording of the query differed from the text. Methods based on semantic similarity, such as FAISS and semantic chunking, performed better than BM25 but still did not reach the performance of SPLADE. The hybrid approach, which combines keyword matching and semantic similarity, generally improved performance when both individual methods worked well. Adding MMR (Maximal Marginal Relevance) to FAISS further improved results by reducing redundancy in the retrieved passages while maintaining relevance. Overall, these results suggest that retrieval methods that understand semantic meaning are more effective than simple keyword matching when searching clinical notes.
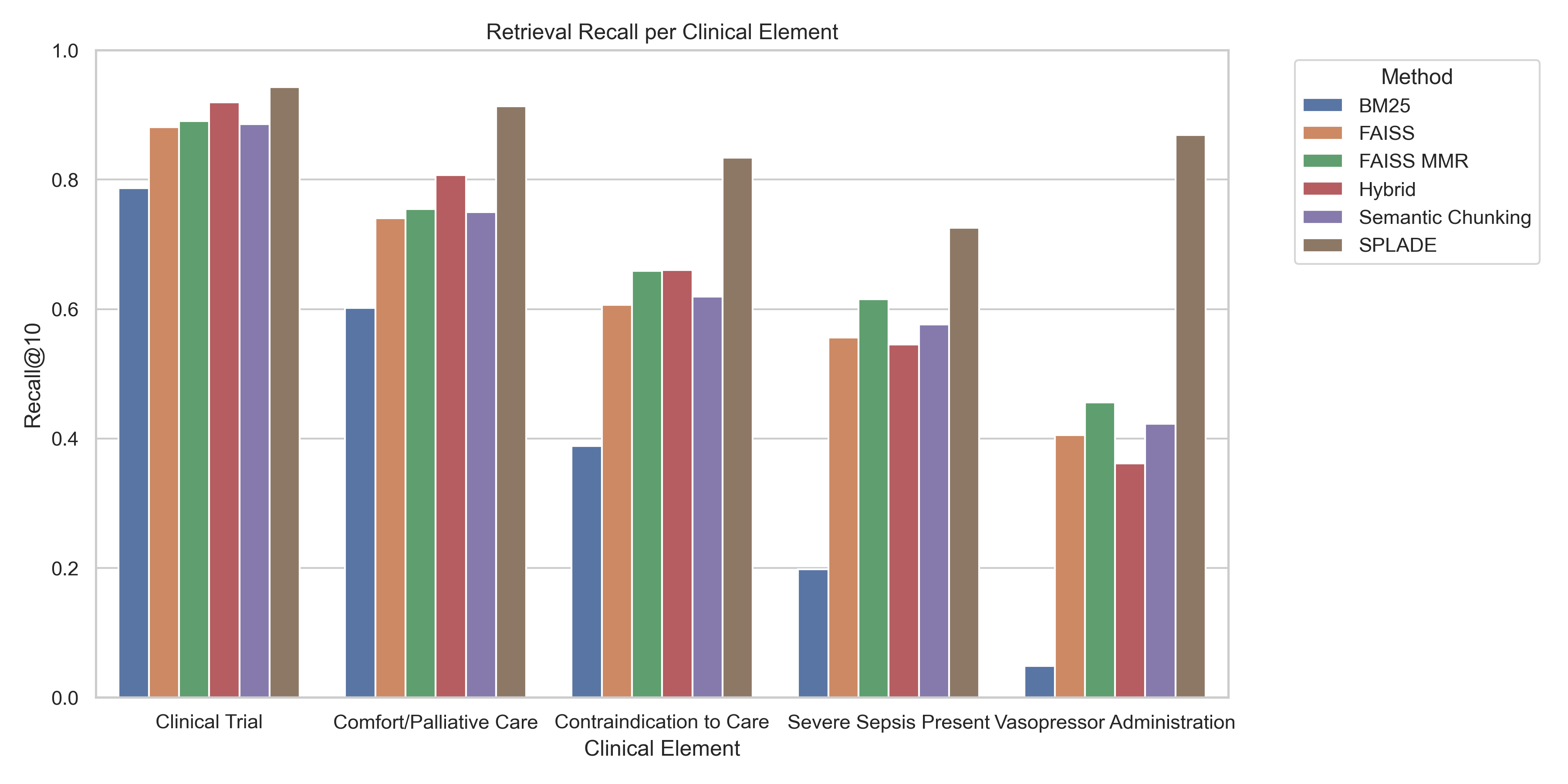
We also found that retrieval success depended on the type of clinical information being searched. Some elements were easier for the LLM to find than others. The easiest elements to retrieve were clinical trial information and comfort care directives. Vasopressor administration was the most difficult to retrieve, with severe sepsis indicators and administrative contraindications to care being moderately difficult. Across nearly all retrieval methods, vasopressor-related information was the hardest to locate, while clinical trial information produced the highest retrieval accuracy. This variation suggests that certain types of clinical language are inherently more difficult for retrieval systems to identify, even when the ground truth evidence is present.
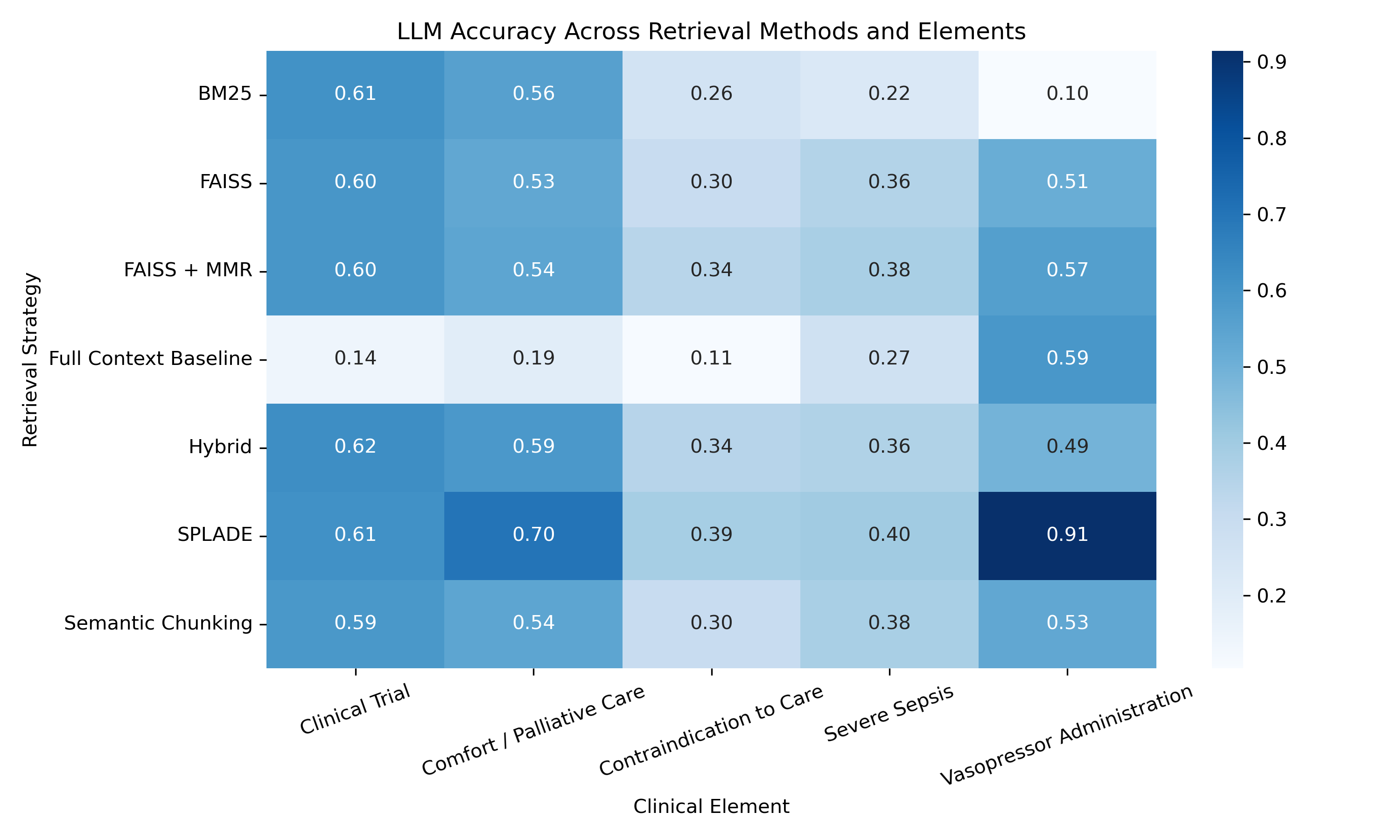
We next evaluated how well the LLM could answer questions and extract evidence from the patient records. When the model received retrieved passages, it generally performed better than when given the entire patient record at once. This shows that focusing the model on relevant sections of text helps improve accuracy. Among all retrieval strategies, SPLADE produced the highest LLM accuracy, FAISS + MMR performed second best, and BM25 consistently produced the lowest accuracy. These results demonstrate that LLMs perform best when they are guided toward the most relevant information rather than processing large, unfiltered documents.
We also studied how the amount of text provided to the model affected performance and found that two key patterns emerged.
- Longer context decreased accuracy, likely because irrelevant information distracted the model.
- Retrieving more passages increased the chance of finding the needle, but too many passages introduced additional noise. Balancing these factors, we found that retrieving the top 10 passages provided the best trade-off between retrieval success and LLM accuracy.
Finally, we examined the relationship between retrieval success and LLM accuracy. We observed a strong positive correlation between the two.
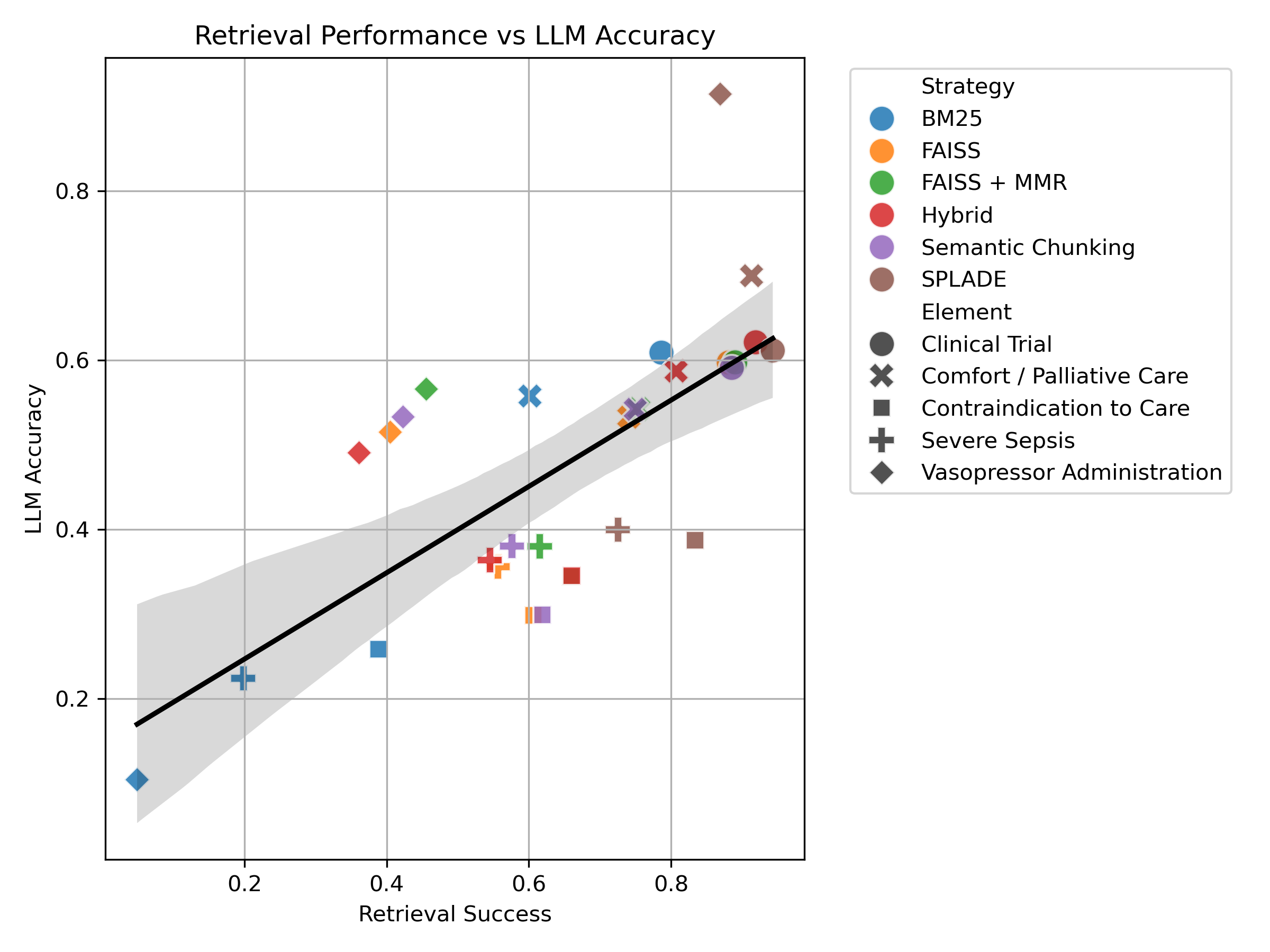
When comparing all the retrieval methods with the baseline, we can see that retrieval methods lead to better LLM accuracy than no retrieval.
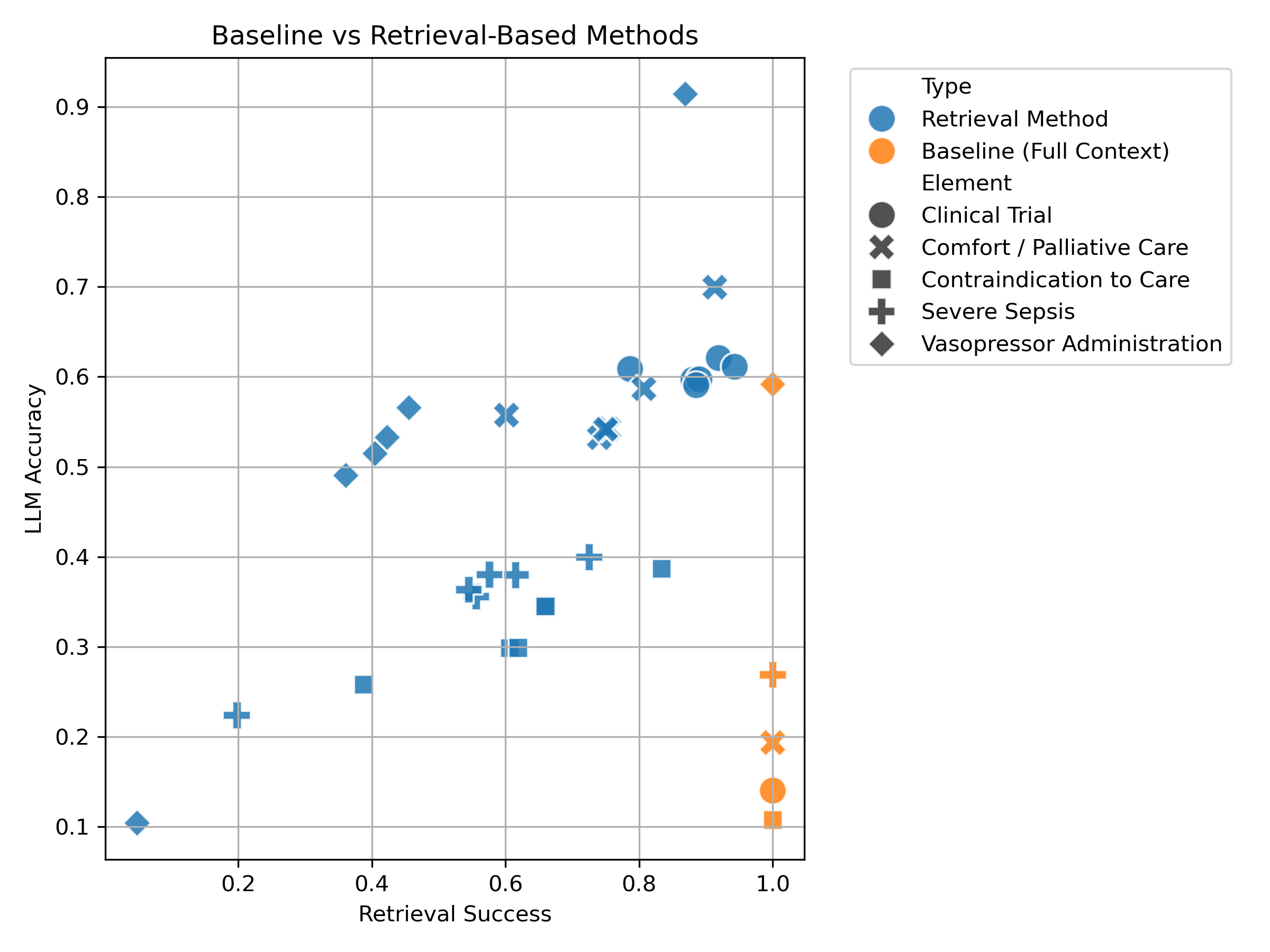
When retrieval methods successfully located the needle, the LLM almost always answered correctly. When retrieval failed to locate the needle, the LLM struggled to produce accurate responses. Interestingly, although the baseline method (providing the full patient record) technically included the needle every time, the LLM performed worse due to the large amount of irrelevant context.
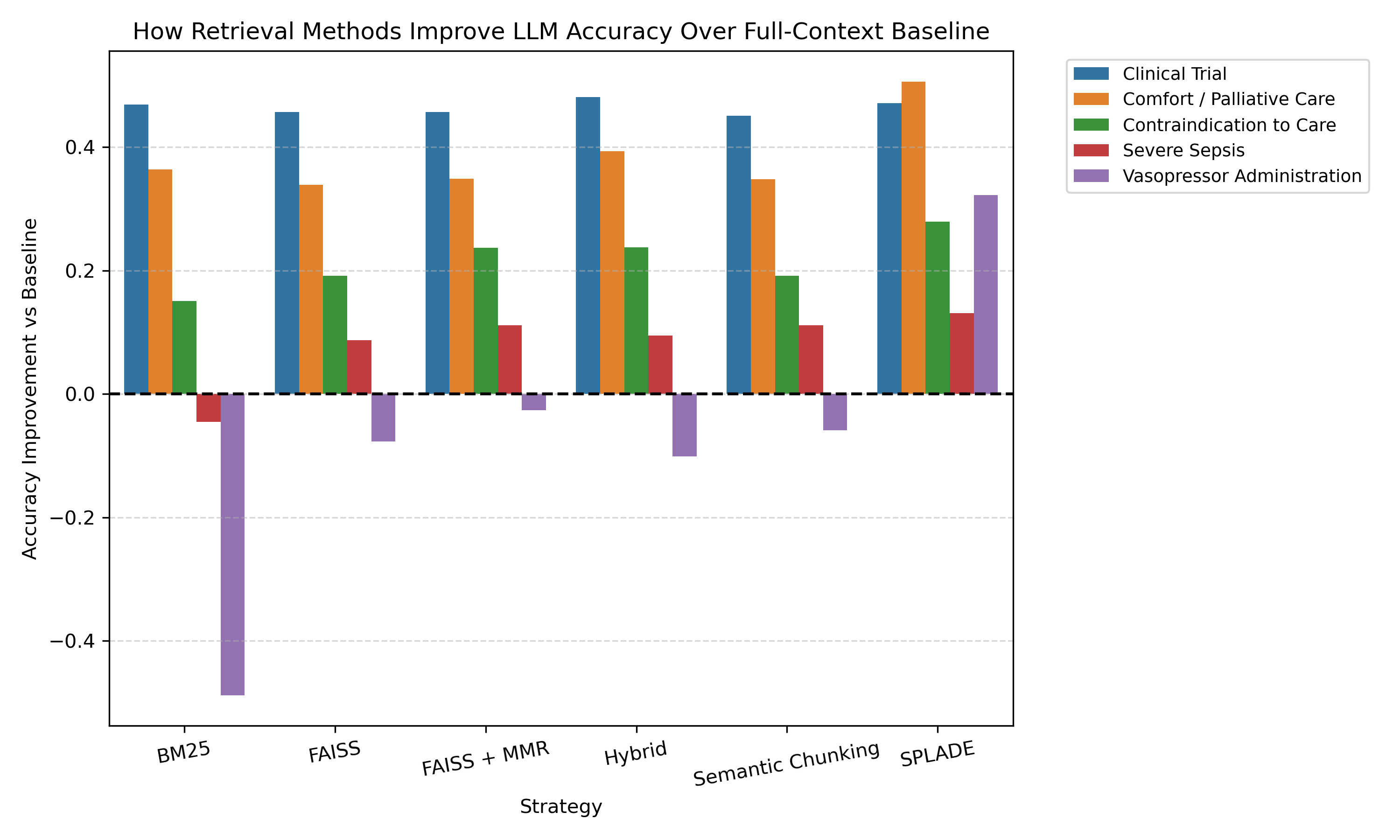
These findings highlight a key takeaway: LLMs are highly effective at extracting clinical information when they are provided with focused high quality context.
Conclusion
Our results show that the success of LLMs in analyzing clinical notes depends heavily on how well relevant information is retrieved. Even powerful models struggle when they are given large unfiltered documents. However, when retrieval methods successfully identify the most relevant passages, LLMs can accurately locate and extract important clinical information.
This finding highlights a key challenge in applying LLMs to healthcare: medical records are long, complex, and often contain distracting information. Improving retrieval strategies can help the LLM focus on the most relevant parts of patient records and avoid errors caused by irrelevant context.
To study this problem safely, we developed a benchmark that allows researchers to evaluate retrieval and extraction methods without exposing sensitive patient data.
Overall, our results suggest that improving retrieval systems may be one of the most effective ways to make AI tools more reliable for analyzing clinical documentation.
Future research will focus on testing these retrieval strategies on more complex clinical datasets and developing improved retrieval methods. As these systems improve, LLMs could become valuable tools for helping clinicians quickly locate critical information within large volumes of medical records, potentially reducing documentation burden and improving access to patient insights.